Desmond_2022-4在4090队列上的表现和批量化MD操作
拳打Amber脚踢Gromacs最终攀上Desmond高峰
说明:
笔者在长期受到Amber、Gromacs模拟系统构建的折磨后,痛定思痛决定下载带有GUI美观、完全自动化、一键分析的Desmond。上海科技大学Wang-Lin搭建了Schrödinger中文社区微信群(公众号:蛋白矿工),实现了使用者的互相交流和问答摘录整理,并就Schrödinger软件的使用编写了大量实用脚本。
基于命令行操作的Amber代码包非常适合生物分子体系的模拟,基于命令行操作的Amber代码包非常适合生物分子体系的模拟,包括AmberTools、Amber两部分。前者免费,但提升生产力的多CPU并行和GPU加速功能需要通过Amber实现,但发展中国家(如中国)可以通过学术申请获得。
同样基于命令行操作的的Gromacs代码包由于上手快、同样基于命令行操作的的Gromacs代码包由于上手快、中文教程详尽等优点收到初学者的一致好评,教程可以参考李继存老师博客(http://jerkwin.github.io/)、B站和计算化学公社(http://bbs.keinsci.com/forum.php)等。 初学者跟着教程做就可以(http://www.mdtutorials.com/),深入学习还是推荐大家去看官方的文档。
初识Desmond是使用世界上最贵的分子模拟软件之一的“初识Desmond是使用世界上最贵的分子模拟软件之一的“学术版”Maestro时,它是由D.E.Shaw Research开发的MD代码,但在Schrödinger公司的帮助下实现了GUI图形界面操作。投资界科学家D.E.Shaw顺手还制造l一台用于分子动力学模拟的专业超级计算机Anton2,当然相对于宇宙最快的Gromacs还是差了点(懂得都懂)。学术和非营利机构用户更是可以在https://www.deshawresearch.com/resources.htmlD.E.Shaw Research官方网站注册信息,免费获得拥有Maestro界面的Desmond软件。
Desmonds的安装:
在D.E.Shaw Research注册后下载相应软件包,上传到HPC集群后解压安装上传到HPC集群个人home下任意位置,随后创建软件文件夹(作为后续安装路径)复制pwd输出的路径内容(笔者为/fsa/home/ljx_zhangzw/software):
mkdir software
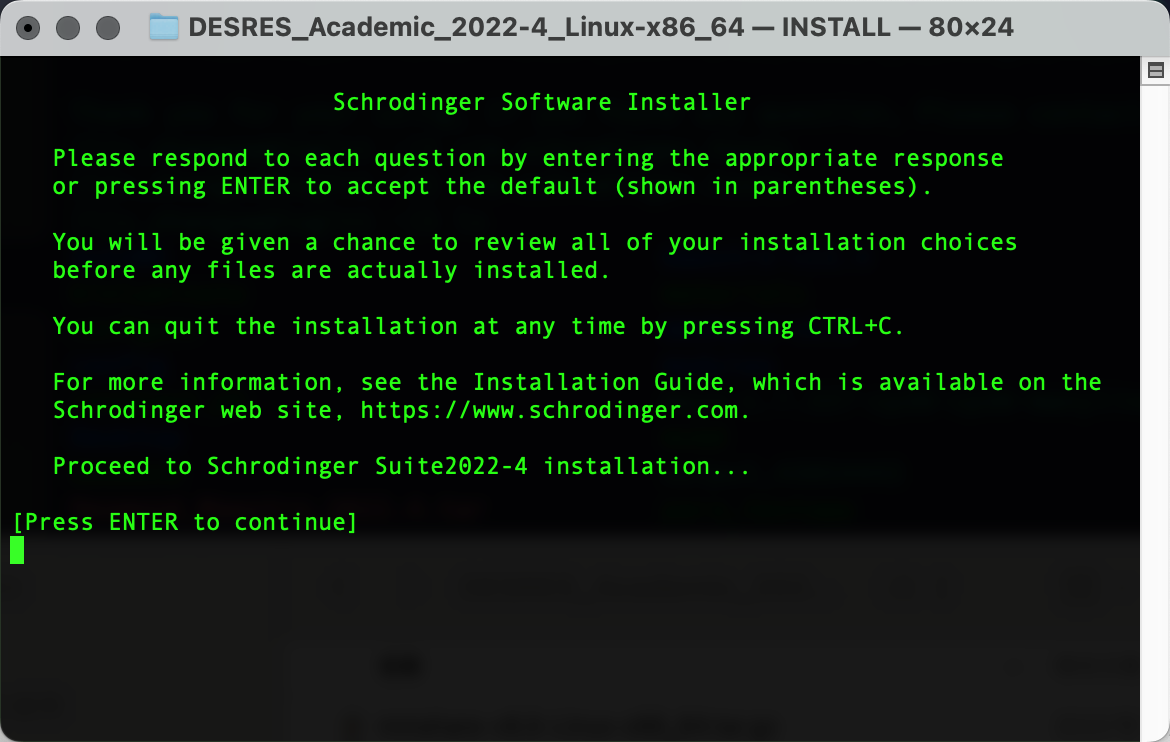
pwd解压安装,进入安装引导界面并按Enter继续进行安装:
tar -xvf Desmond_Maestro_2022.4.tar
cd DESRES_Academic_2022-4_Linux-x86_64
./INSTALL在D.E.Shaw Research注册后下载相应软件包,上传到HPC集群后解压安装
文件位置:
/fs00/software/singularity-images/ngc\_gromacs\_2021.3.sif
提交代码:
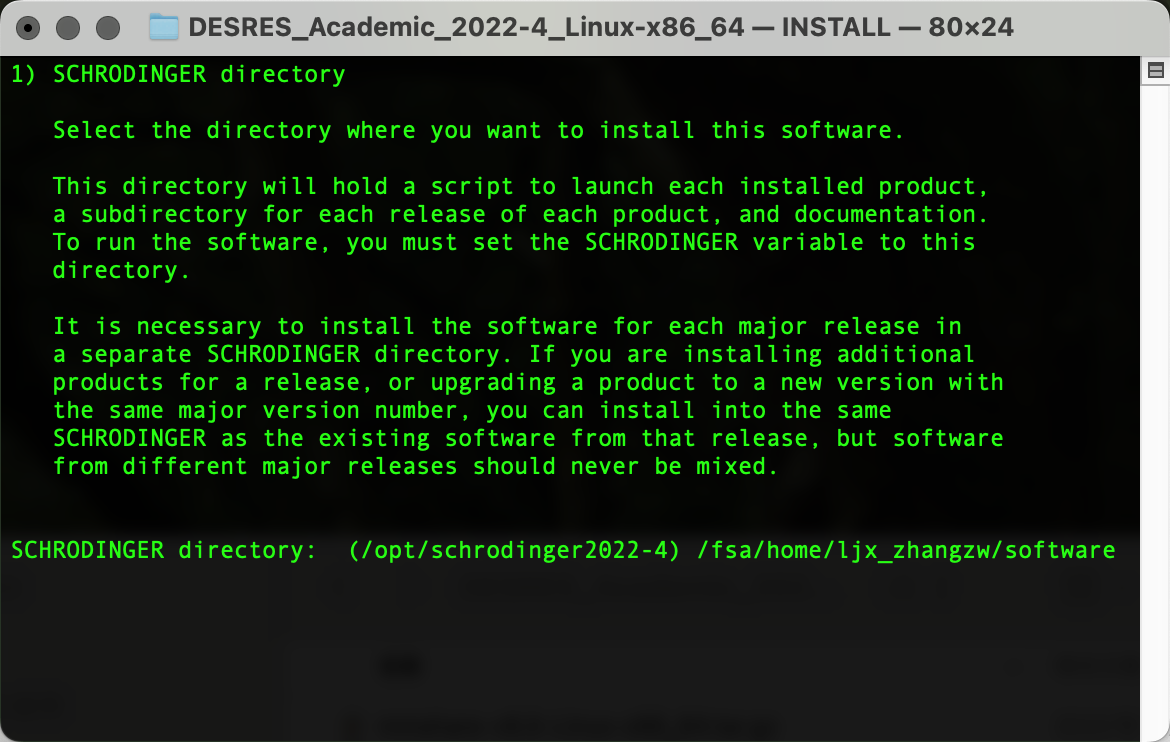
输出以下界面后粘贴复制pwd输出的路径内容,Enter确认为安装路径:
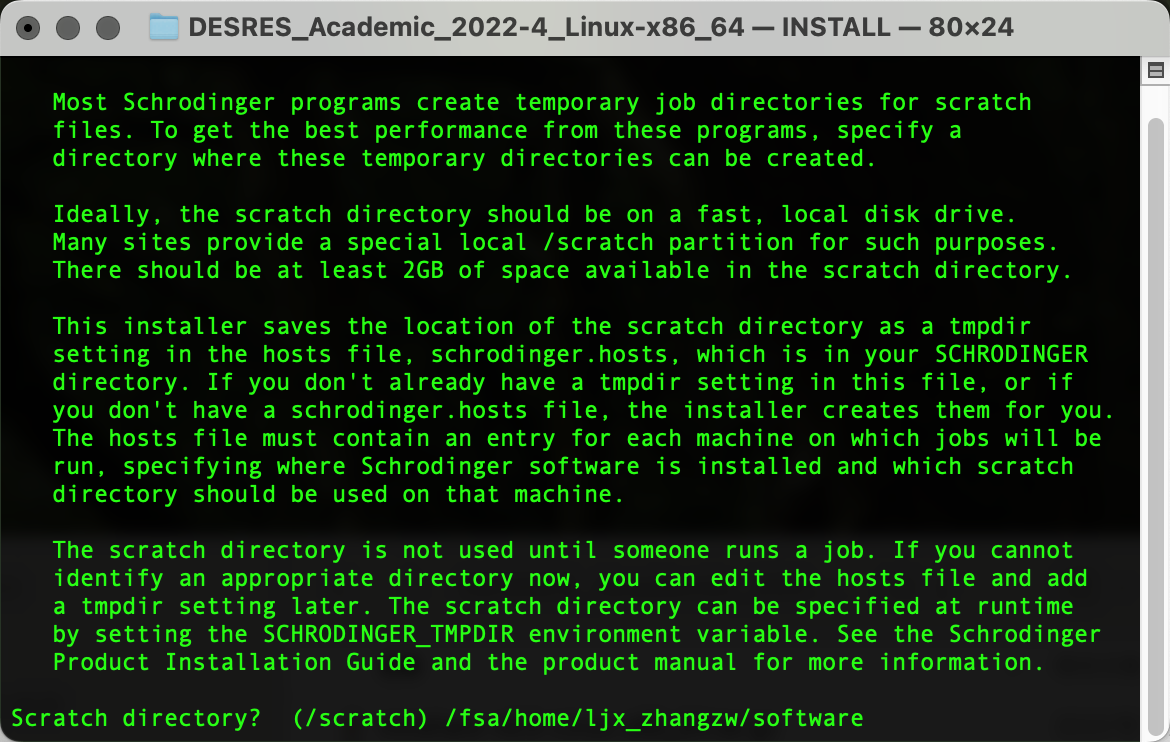
输出以下界面后粘贴复制pwd输出的路径内容,Enter确认为临时文件路径:
输出以下界面后输入y,Enter确认上述两个路径并进行安装:
安装完成后进入安装路径,并写入环境变量:
#BSUBcd /fsa/home/ljx_zhangzw/software
echo "export Desmond=${PWD}/" >> ~/.bashrc同时由于Desmond对集群计算中提交任务队列的要求,需要在安装路径修改schrodinger.host以定义GPU信息,示例给出的是83a100ib队列中m001节点信息:
#localhost
name:localhost
temdir:/fsa/home/ljx_zhangzw/software
#HPC
name: HPC
host: m001
queue: LSF
qargs: -q 72rtxib
#BSUB83a100ib -gpu "num=1"1
module load singularity/latest
export OMP_NUM_THREADS="$LSB_DJOB_NUMPROC"
SINGULARITY="singularity run --nvtmpdir: /fs00/software/singularity-images/ngc_gromacs_2021.3.sif"fsa/home/ljx_zhangzw/software
${SINGULARITY}schrodinger: gmx/fsa/home/ljx_zhangzw/software
gromppgpgpu: -f0, minim.mdpNVIDIA -cA100
1aki_solv_ions.grogpgpu: -p1, topol.topNVIDIA -oA100
em.tprgpgpu: ${SINGULARITY}2, gmxNVIDIA mdrunA100
-nbgpgpu: gpu3, -ntmpiNVIDIA 2A100
-deffnmgpgpu: em4, NVIDIA A100
gpgpu: 5, NVIDIA A100
gpgpu: 6, NVIDIA A100
gpgpu: 7, NVIDIA A100平衡模拟(nvt)
#BSUB -q 72rtxib
#BSUB -gpu "num=1"
module load singularity/latest
export OMP_NUM_THREADS="$LSB_DJOB_NUMPROC"
SINGULARITY="singularity run --nv /fs00/software/singularity-images/ngc_gromacs_2021.3.sif"
${SINGULARITY} gmx grompp -f nvt.mdp -c em.gro -r em.gro -p topol.top -o nvt.tpr
${SINGULARITY} gmx mdrun -nb gpu -ntmpi 2 -deffnm nvt
平衡模拟(npt)
#BSUB -q 72rtxib
#BSUB -gpu "num=1"
module load singularity/latest
export OMP_NUM_THREADS="$LSB_DJOB_NUMPROC"
SINGULARITY="singularity run --nv /fs00/software/singularity-images/ngc_gromacs_2021.3.sif"
${SINGULARITY} gmx grompp -f npt.mdp -c nvt.gro -r nvt.gro -t nvt.cpt -p topol.top -o npt.tpr
${SINGULARITY} gmx mdrun -nb gpu -ntmpi 2 -deffnm npt
成品模拟(md)
#BSUB -q 723090ib
#BSUB -gpu "num=1"
module load singularity/latest
export OMP_NUM_THREADS="$LSB_DJOB_NUMPROC"
SINGULARITY="singularity run --nv /fs00/software/singularity-images/ngc_gromacs_2021.3.sif"
${SINGULARITY} gmx grompp -f md.mdp -c npt.gro -t npt.cpt -p topol.top -o md_0_1.tpr
${SINGULARITY} gmx mdrun -nb gpu -bonded gpu -update gpu -pme gpu -pmefft gpu -deffnm md_0_1
成品模拟(md)需要注意的是:
也可以参照以下命令进行修改,以作业脚本形式进行提交:1、修改tmpdir和schrodinger对应路径为自己的安装路径;
#BSUB2、节点host、队列83a100ib以及显卡信息gpgpu: -q0-7, 723090ibNVIDIA #BSUB -gpu "num=1"
module load singularity/latest
export OMP_NUM_THREADS="$LSB_DJOB_NUMPROC"
SINGULARITY="singularity run --nv /fs00/software/singularity-images/ngc_gromacs_2021.3.sif"
${SINGULARITY} gmx pdb2gmx -f protein.pdb -o protein_processed.gro -water tip3p -ignh -merge all <<< 4
${SINGULARITY} gmx editconf -f protein_processed.gro -o pro_newbox.gro -c -d 1.0 -bt cubic
${SINGULARITY} gmx solvate -cp pro_newbox.gro -cs spc216.gro -o pro_solv.gro -p topol.top
${SINGULARITY} gmx grompp -f ../MDP/ions.mdp -c pro_solv.gro -p topol.top -o ions.tpr
${SINGULARITY} gmx genion -s ions.tpr -o pro_solv_ions.gro -p topol.top -pname NA -nname CL -neutral <<< 13
${SINGULARITY} gmx grompp -f ../MDP/minim.mdp -c pro_solv_ions.gro -p topol.top -o em.tpr
${SINGULARITY} gmx mdrun -v -deffnm em
${SINGULARITY} gmx energy -f em.edr -o potential.xvg <<< "10 0"
${SINGULARITY} gmx grompp -f ../MDP/nvt.mdp -c em.gro -r em.gro -p topol.top -o nvt.tpr
${SINGULARITY} gmx mdrun -deffnm nvt
${SINGULARITY} gmx energy -f nvt.edr -o temperature.xvg <<< "16 0"
${SINGULARITY} gmx grompp -f ../MDP/npt.mdp -c nvt.gro -r nvt.gro -t nvt.cpt -p topol.top -o npt.tpr
${SINGULARITY} gmx mdrun -deffnm npt
${SINGULARITY} gmx energy -f npt.edr -o pressure.xvg <<< "18 0"
${SINGULARITY} gmx grompp -f ../MDP/md.mdp -c npt.gro -t npt.cpt -p topol.top -o md.tpr
${SINGULARITY} gmx mdrun -v -deffnm md
${SINGULARITY} gmx rms -f md.xtc -s md.tpr -o rmsd.xvg <<< "4 4"
软件信息:
GROMACS version: 2021.3-dev-20210818-11266ae-dirty-unknown
Precision: mixed
Memory model: 64 bit
MPI library: thread_mpi
OpenMP support: enabled (GMX_OPENMP_MAX_THREADS = 64)
GPU support: CUDA
SIMD instructions: AVX2_256
FFT library: fftw-3.3.9-sse2-avx-avx2-avx2_128-avx512
CUDA driver: 11.20
CUDA runtime: 11.40
测试算例:
ATOM 218234 (401 Protein residues, 68414 SOL, 9 Ion residues)
nsteps = 100000000 ; 200 ns
eScience中心GPU测试:3、对于不同的任务调度系统,Schrödinger公司KNOWLEDGE 能量最小化(em)、平衡模拟(nvt、npt)使用1个GPU进行模拟,成品模拟(md)使用1个GPU进行模拟。BASE和蛋白矿工知乎号Q2进行了介绍;
4、截止2023年6月16日,HPC集群可供Desmond使用的计算卡(而非3090、4090等)包括:
2*NVIDIA Tesla K40 12GB
128GB RAM
56Gb FDR InfiniBand
x001
gpgpu: 0, NVIDIA Tesla K40
gpgpu: 1, NVIDIA Tesla K40
校内及协同创新中心用户
1.2 元/卡/小时=0.1元/核/小时
1.2 =0.12*NVIDIA Tesla P100 PCIe 16GB
128GB RAM
56Gb FDR InfiniBand
x002
gpgpu: 0, NVIDIA Tesla P100
gpgpu: 1, NVIDIA Tesla P100
校内及协同创新中心用户
1.68 元/卡/小时=0.12元/核/小时
8*NVIDIA Tesla V100 SXM2 32GB
768GB RAM
100Gb EDR InfiniBand
n002
gpgpu: 0, NVIDIA Tesla V100
gpgpu: 1, NVIDIA Tesla V100
gpgpu: 2, NVIDIA Tesla V100
gpgpu: 3, NVIDIA Tesla V100
gpgpu: 4, NVIDIA Tesla V100
gpgpu: 5, NVIDIA Tesla V100
gpgpu: 6, NVIDIA Tesla V100
gpgpu: 7, NVIDIA Tesla V100
校内及协同创新中心用户
3 元/卡/小时=0.6元/核/小时
72rtxib AMD EPYC 7302
4*NVIDIA TITAN RTX 24GB
128GB RAM
100Gb HDR100 InfiniBand
g005 or g006 or g007
gpgpu: 0, NVIDIA TITAN RTX
gpgpu: 1, NVIDIA TITAN RTX
gpgpu: 2, NVIDIA TITAN RTX
gpgpu: 3, NVIDIA TITAN RTX
校内及协同创新中心用户
1.8 元/卡/小时=0.45元/核/小时
8*NVIDIA Tesla A100
512GB
200Gb
m001
gpgpu:
0,gpgpu:
gpgpu:
gpgpu:
gpgpu:
gpgpu: 6, NVIDIA A100
gpgpu: 7, NVIDIA A100
校内及协同创新中心用户
4.8 元/卡/小时=0.6元/核/小时
结论:
单次MD:
能量最小化(em)在任务较少的722080tiib和72rtxib队列中,Run time分别为88.83 ± 12.45和83.25 ± 11.44s;在个人笔记本(Linux)上依照上述步骤安装,或使用Windows浏览器以图形界面Web登陆HPC集群进行点击操作。
教程可参考2022薛定谔中文网络培训和知乎内容。
批量MD:
平衡模拟(nvt)任务在722080tiib、72rtxib和723090ib队列中,Run1、plmd/DSMDrun: time分别为1776.50 ± 181.73、357.00 ± 4.08和309.50 ± 39.06 s;Desmond分子动力学模拟一键式运行脚本:
平衡模拟(npt)任务在722080tiib、72rtxib和723090ib队列中,Run脚本介绍:Wang-Lin-boop/Schrodinger-Script: time分别为5411.00Some ±scripts 247.49、371.25to ±run 6.08和336.50Schrödinger ±jobs 16.68on s;HPC or localhost. (github.com)
通过以上脚本实现当前路径下所有的mae文件(类似于PDB的结构文件,需通过Maestro转存)按照设置参数进行模拟,plmd
-h中给出了示例和详细的输入命令介绍:原子数218234的
Usage: plmd [OPTION] <parameter>
An automatic Desmond MD pipline for protein-ligand complex MD simulation.
Example:
1) plmd -i "*.mae" -S INC -P "chain.name A" -L "res.ptype UNK" -H HPC_CPU -G HPC_GPU
2) plmd -i "*.mae" -S OUC -P "chain.name A" -L "chain.name B" -t 200 ns成品模拟(md)任务在722080tiib、72rtxib、和723090ib队列中,-H HPC_CPU -G HPC_gpu01
3) plmd -i "*.mae" -S "TIP4P:Cl:0.15-Na-Cl+0.02-Fe2-Cl+0.02-Mg2-Cl" -L "res.num 999" -G HPC_gpu03
4) plmd -i "*.cms" -P "chain.name A" -L "res.ptype ADP" -H HPC_CPU -G HPC_gpu04
Input parameter:
-i Use a file name (Multiple files are wrapped in "", and split by ' ') *.mae or *.cms ;
or regular expression to represent your input file, default is *.mae.
System Builder parameter:
-S System Build Mode: <INC>
INC: System in cell, salt buffer is 0.15M KCl, water is TIP3P. Add K to neutralize system.
OUC: System out of cell, salt buffer is 0.15M NaCl, water is TIP3P. Add Na to neutralize system.
Custom Instruct: Such as: "TIP4P:Cl:0.15-Na-Cl+0.02-Fe2-Cl+0.02-Mg2-Cl"
Interactive addition of salt. Add Cl to neutralize system.
for positive_ion: Na, Li, K, Rb, Cs, Fe2, Fe3, Mg2, Ca2, Zn2 are predefined.
for nagative_ion: F, Cl, Br, I are predefined.
for water: SPC, TIP3P, TIP4P, TIP5P, DMSO, METHANOL are predefined.
-b Define a boxshape for your systems. <cubic>
box types: dodecahedron_hexagon, cubic, orthorhombic, triclinic
-s Define a boxsize for your systems. <15.0>
for dodecahedron_hexagon and cubic, defulat is 15.0;
for orthorhombic or triclinic box, defulat is [15.0 15.0 15.0];
If you want use Orthorhombic or Triclinic box, your parameter should be like "15.0 15.0 15.0"
-R Redistribute the mass of heavy atoms to bonded hydrogen atoms to slow-down high frequency motions.
-F Define a force field to build your systems. <OPLS_2005>
OPLS_2005, S-OPLS, OPLS3e, OPLS3, OPLS2 are recommended to protein-ligand systems.
Simulation control parameter:
-m Enter the maximum simulation time for the Brownian motion simulation, in ps. <100>
-t Enter the Molecular dynamics simulation time for the product simulation, in ns. <100>
-T Specify the temperature to be used, in kelvin. <310>
-N Number of Repeat simulation with different random numbers. <1>
-P Define a ASL to protein, such as "protein".
-L Define a ASL to ligand, such as "res.ptype UNK".
-q Turn off protein-ligand analysis.
-u Turn off md simulation, only system build.
-C Set constraint to an ASL, such as "chain.name A AND backbone"
-f Set constraint force, default is 10.
-o Specify the approximate number of frames in the trajectory. <1000>
This value is coupled with the recording interval for the trajectory and the simulation time: the number of frames times the trajectory recording interval is the total simulation time.
If you adjust the number of frames, the recording interval will be modified.
Job control:
-G HOST of GPU queue, default is HPC_GPU.
-H HOST of CPU queue, default is HPC_CPU.
-D Your Desmond path. <$Desmond>性能表现差别不大2、AutoMD: Desmond分子动力学模拟一键式运行脚本:,分别为110.03 ± 2.75、115.83 ± 1.54和107.66 ± 5.90 ns/day。
下载地址:Wang-Lin-boop/AutoMD: Easy to get started with molecular dynamics simulation. (github.com)
综上,建议在能量最小化(em)、平衡模拟(nvt、npt)等阶段使用排队任务较少的72rtxib队列脚本介绍:AutoMD:从初始结构到MD轨迹,只要一行Linux命令? ,建议在成品模拟(md)阶段按照任务数量(从笔者使用情况来看,排队任务数量72rtxib<722080tiib<723090ib<83a100ib)、GPU收费情况(校内及协同创新中心用户:72rtxib队列1.8- 元/卡/小时=0.45元/核/小时、722080tiib队列1.2知乎 元/卡/小时=0.3元/核/小时、723090ib队列1.8 元/卡/小时=0.3元/核/小时、83a100ib队列4.8 元/卡/小时=0.3元/核/小时)适当考虑队列。(zhihu.com)
由于Desmond学术版本提供的力场有限,plmd无法有效引入其他力场参数,因此在plmd的迭代版本AutoMD上,通过配置D.E.Shaw
Research开发的Viparr和Msys代码在Desmond中引入如Amber,Charmm等力场来帮助模拟系统的构建。在以上提交代码中,未涉及到Gromacs的并行效率问题(直接“num=4”并不能在集群同时使用4块GPU#在HPC上安装Viparr和Msys),感兴趣的同学可以查看http://bbs.keinsci.com/thread-13861-1-1.html以及https://developer.nvidia.com/blog/creating-faster-molecular-dynamics-simulations-with-gromacs-2020/的相关解释。但根据前辈的经验,ATOM 500000以上才值得使用两张GPU加速卡,原因在于Gromacs的并行效率不明显。感兴趣的同学也可以使用Amber的GPU并行加速,但对显卡的要求为3090或者tesla A100。这里提供了GPU=4的gromacs命令:
在个人笔记本(Linux)上下载以下代码文件,并上传至集群个人home的software中:
gmxwget mdrunhttps://github.com/DEShawResearch/viparr/releases/download/4.7.35/viparr-4.7.35-cp38-cp38-manylinux2014_x86_64.whl
wget https://github.com/DEShawResearch/msys/releases/download/1.7.337/msys-1.7.337-cp38-cp38-manylinux2014_x86_64.whl
git clone git://github.com/DEShawResearch/viparr-ffpublic.git
git clone https://github.com/Wang-Lin-boop/AutoMD随后进入desmond工作目录,启动虚拟环境以帮助安装Viparr和Msys:
cd /fsa/home/ljx_zhangzw/software
./run schrodinger_virtualenv.py schrodinger.ve
source schrodinger.ve/bin/activate
pip install -deffnm-upgrade pip
pip install msys-1.7.337-cp38-cp38-manylinux2014_x86_64.whl
pip install viparr-4.7.35-cp38-cp38-manylinux2014_x86_64.whl
echo "export viparr=${PWD}/schrodinger.ve/bin" >> ~/.bashrc同时,将viparr-ffpublic.git解压并添加到环境变量:
echo "export VIPARR_FFPATH=${PWD}/viparr-ffpublic/ff" >> ~/.bashrc最后,将AutoMD.git解压并进行安装,同时指定可自动添加补充力场:
cd AutoMD
echo "alias AutoMD=${PWD}/AutoMD" >> ~/.bashrc
chmod +x AutoMD
source ~/.bashrc
cp -r ff/* ${VIPARR_FFPATH}/#AutoMD -h
在输入命令介绍中,给出了四个示例:胞质蛋白-配体复合物、血浆蛋白-蛋白复合物、DNA/RNA-蛋白质复合物、以及需要Meastro预准备的膜蛋白。在这些示例中:
通过-i命令输入了当前路径所有的复合物结构.mae文件(也可以是Maestro构建好的模拟系统.cms文件);
通过-S指定了模拟系统的溶液环境,包括INC(0.15M KCl、SPC水、K+中和)、OUC(0.15M NaCl、SPC水、Na+中和),以及自定义溶液环境如"SPC:Cl:0.15-K-Cl+0.02-Mg2-Cl";
通过-b和-s定义了模拟系统的Box形状和大小;
通过-F定义力场参数,由于学术版Desmond的要求不被允许使用S-OPLS力场,仅可使用OPLS_2005或其他力场;OPLS适用于蛋白-配体体系,Amber适用于蛋白-核酸,Charmm适用于膜蛋白体系,DES-Amber适用于PPI复合体体系,但是这些搭配并不是绝对的,我们当然也可以尝试在膜蛋白体系上使用Amber力场,在核酸体系上使用Charmm力场,具体的情况需要用户在自己的体系上进行尝试;
Usage: AutoMD [OPTION] <parameter>
Example:
1) MD for cytoplasmic protein-ligand complex:
AutoMD -i "*.mae" -S INC -P "chain.name A" -L "res.ptype UNK" -F "S-OPLS"
2) MD for plasma protein-protein complex:
AutoMD -i "*.mae" -S OUC -F "DES-Amber"
3) MD for DNA/RNA-protein complex:
AutoMD -i "*.mae" -S "SPC:Cl:0.15-K-Cl+0.02-Mg2-Cl" -F Amber
4) MD for membrane protein, need to prior place membrane in Meastro.
AutoMD -i "*.mae" -S OUC -l "POPC" -r "Membrane" -F "Charmm"
Input parameter:
-i Use a file name (Multiple files are wrapped in "", and split by ' ') *.mae or *.cms ;
or regular expression to represent your input file, default is *.mae.
System Builder parameter:
-S System Build Mode: <INC>
INC: System in cell, salt buffer is 0.15M KCl, water is SPC. Add K to neutralize system.
OUC: System out of cell, salt buffer is 0.15M NaCl, water is SPC. Add Na to neutralize system.
Custom Instruct: Such as: "TIP4P:Cl:0.15-Na-Cl+0.02-Fe2-Cl+0.02-Mg2-Cl"
Interactive addition of salt. Add Cl to neutralize system.
for positive_ion: Na, Li, K, Rb, Cs, Fe2, Fe3, Mg2, Ca2, Zn2 are predefined.
for nagative_ion: F, Cl, Br, I are predefined.
for water: SPC, TIP3P, TIP4P, TIP5P, DMSO, METHANOL are predefined.
-l Lipid type for membrane box. Use this option will build membrane box. <None>
Lipid types: POPC, POPE, DPPC, DMPC.
-b Define a boxshape for your systems. <cubic>
box types: dodecahedron_hexagon, cubic, orthorhombic, triclinic
-s Define a boxsize for your systems. <15.0>
for dodecahedron_hexagon and cubic, defulat is 15.0;
for orthorhombic or triclinic box, defulat is [15.0 15.0 15.0];
If you want use Orthorhombic or Triclinic box, your parameter should be like "15.0 15.0 15.0"
-R Redistribute the mass of heavy atoms to bonded hydrogen atoms to slow-down high frequency motions.
-F Define a force field to build your systems. <OPLS_2005>
OPLS_2005, S-OPLS are recommended to receptor-ligand systems.
Amber, Charmm, DES-Amber are recommended to other systems. Use -O to show more details.
Use the "Custom" to load parameters from input .cms file.pdb.
Simulation control parameter:
-m Enter the maximum simulation time for the Brownian motion simulation, in ps. <100>
-r The relaxation protocol before MD, "Membrane" or "Solute". <Solute>
-e The ensemble class in MD stage, "NPT", "NVT", "NPgT". <NPT>
-t Enter the Molecular dynamics simulation time for the product simulation, in ns. <100>
-T Specify the temperature to be used, in kelvin. <310>
-N Number of Repeat simulation with different random numbers. <1>
-P Define a ASL to receptor, such as "protein".
-L Define a ASL to ligand and run interaction analysis, such as "res.ptype UNK".
-u Turn off md -ntmpisimulation, 4only system build.
-ntompC 7Set constraint to an ASL, such as "chain.name A AND backbone"
-npmef 1Set constraint force, default is 10.
-nbo gpuSpecify the approximate number of frames in the trajectory. <1000>
This value is coupled with the recording interval for the trajectory and the simulation time:
the number of frames times the trajectory recording interval is the total simulation time.
If you adjust the number of frames, the recording interval will be modified.
Job control:
-pmeG gpuHOST of GPU queue, default is GPU.
-bondedH gpuHOST of CPU queue, default is CPU.
-pmefftD gpuYour Desmond path. </fsa/home/ljx_zhangzw/>
-V Your viparr path. </fsa/home/ljx_zhangzw/schrodinger.ve/bin>
-v Your viparr force fields path. </fsa/home/ljx_zhangzw/viparr-ffpublic/ff>示例任务提交脚本及命令:
#BSUB -L /bin/bash
#BSUB -q 734090ib
module load cuda/12.0.0
source /fsa/home/ljx_zhangzw/.bashrc
echo $PATH
echo $LD_LIBRARY_PATH
AutoMD -i "*.mae" -S OUC -P "chain.name A" -L "res.ptype UNK" -F "OPLS_2005" -T 298 -s 10 -H m006 -G m006在模拟参数控制命令中,通过-P和-L分别定义了mae文件中的受体和配体,-m默认
-T定义体系温度为298 K,默认进行100 ns和1次重复模拟次数,通过-H和-G指定了host文件中的相应计算队列。
性能表现待更新....